Happy fourth birthday, DiSSCover!
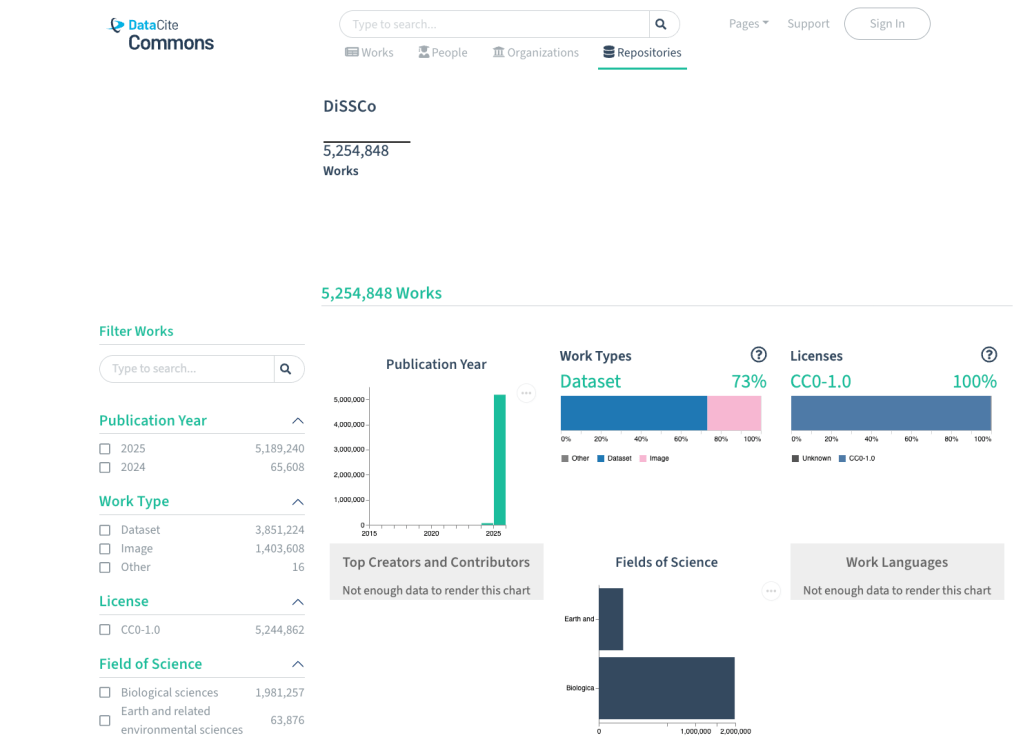
Four years ago, DiSSCover was born with an ambitious mission: to play an active role in driving a digital revolution in collections research by digitising European specimen collections and working towards enriched digital specimen data through expert annotations. The application started out as a Proof of Concept, or in other words, a way to showcase the many functionalities and features that DiSSCo has to offer, like allowing users to search through millions of records of collection data, add expert annotations and collaborate on FAIR data.
In those four years, our DiSSCo ecosystem has grown rapidly. While DiSSCover has always been about the digitisation of the natural history collections of more than 23 European countries, it has also been about accelerating science. From machine annotation services that translate original labels to AI that identifies specimens in a single image, DiSSCover has outgrown the status of ‘Proof of Concept’ and needs to go through puberty. We need a platform built to last.
Maturing DiSSCover
With an application as extensive as DiSSCover 1.0, the technical debt became too big. The application simply became too large to maintain efficiently; bugs became harder to squash; implementing features required unnecessarily complex solutions. On top of all that, our users’ needs had changed. Or – to be more precise – we onboarded actual users who provided us with very valuable feedback.
We took a hard look at where we were and where we wanted to be. Then, with the help of our UX designer, Martijn de Visser, and direct feedback from our community, we decided to strip away what wasn’t working and rewrite the future of DiSSCover.
The 5 pillars of DiSSCover 2.0
On the one hand, we had to solve the issues described above. On the other hand, this provided us with a great opportunity to implement other features of the application that have been waiting for a long time, or are mandated by law now. So we took this chance to rethink not only the design, but also the implementation. The results of this effort are below:
User-centric design: We’ve created a tight feedback loop with our key user groups. By discussing flows and testing features early, we ensure the application works the way our users want.
True mobility: Whether you are at a desktop, using a tablet in the field, or (theoretically) checking data on your smart fridge, DiSSCover will be fully responsive.
Accessibility first: We believe science belongs to everyone. We are adhering to WCAG AA standards, ensuring that the majority of people living with disabilities can navigate our data without barriers.
A modern tech stack: Built with React, TypeScript, and Web Components, the new DiSSCover is modular and reusable. This means faster updates and a much more stable experience.Digital sustainability: We are mindful of our digital footprint. By optimizing API calls, improving caching, and limiting application weight, we’re making the platform as “green” as the current tech allows.
The agile rollout
So you might be wondering when you can see and enjoy the new DiSSCover 2.0. Well, you can right now! We are adopting the agile approach of deploying small, functional batches.
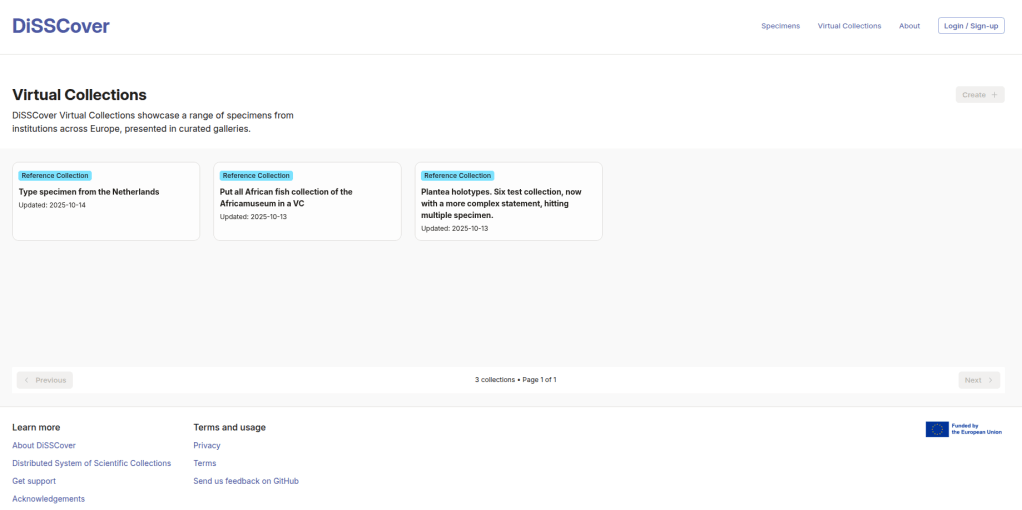
Since January, we’ve released changes to the application. If you head over to disscover.dissco.eu and click on Virtual Collections, you’ll see a completely redesigned experience. Over the coming months, more pages will be updated until the entire platform is refreshed and ready for the future.
Coming Up Next: > How do you actually build for users? In our next post, we’ll dive deep into our User Group sessions and how direct community feedback is shaping our development process. Stay tuned!